Q&A Feature: How 4basebio’s master template generation step shapes next-generation DNA
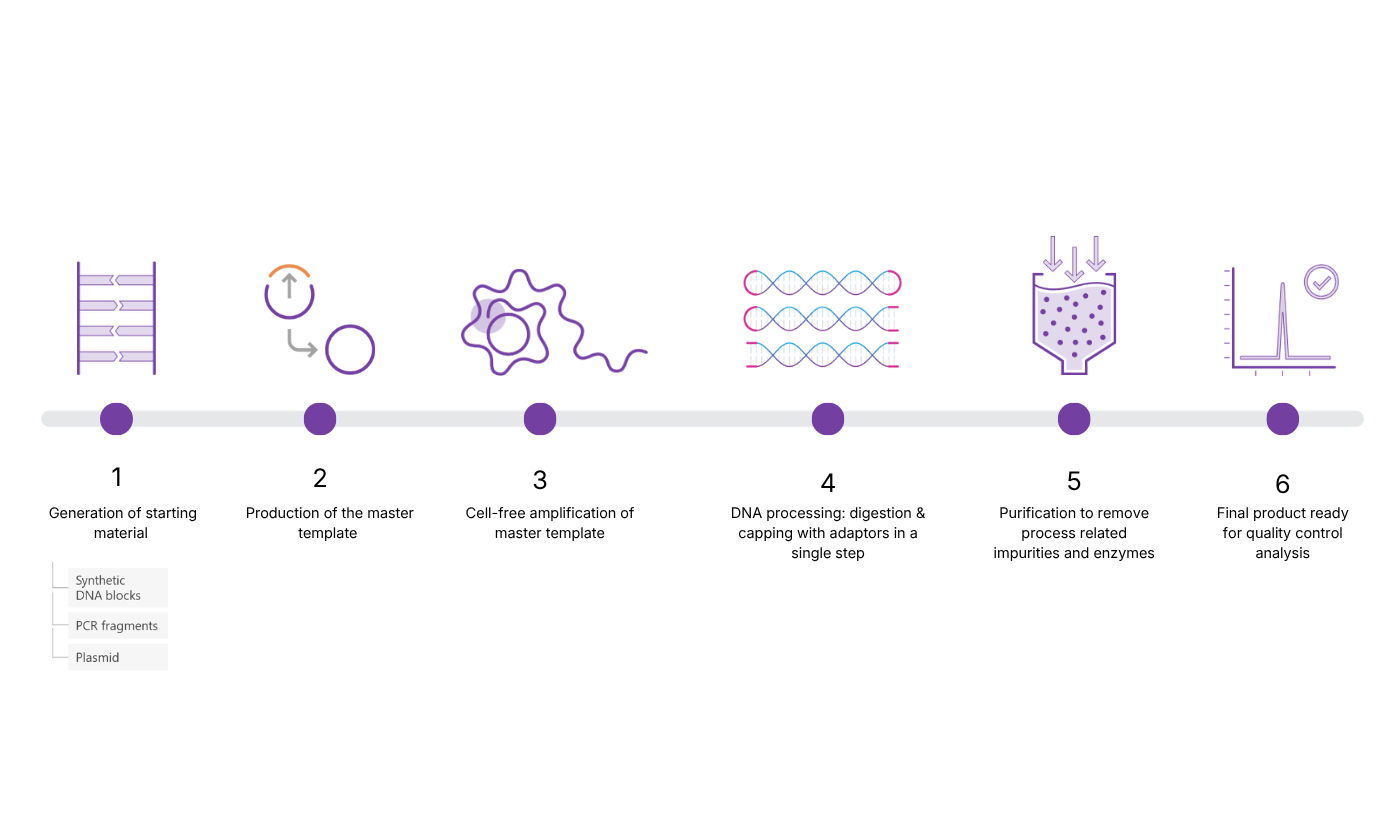
A key step in 4basebio’s synthetic DNA manufacturing process is the master template generation step. This step is unique to our platform and a key differentiator, as it provides control over DNA quality, ensures consistent manufacturing, and improves performance and safety in downstream applications like AAV and mRNA therapeutics.
In this Q&A feature, Tejaswini Mahadeva, Template Generation Team Lead at 4basebio discusses how this critical step works, why it matters for reliable high-quality DNA, and what it means for the future of genetic medicine manufacturing.
Q: What is the master template generation step, and why does this step fundamentally shape the quality and performance of the final synthetic DNA product?
Tejaswini: The template generation step at 4basebio is vital, it involves circularization of linear DNA fragment(s) that will make up the target DNA sequence. The linear DNA fragments can be sources from different origins as required. These can include precursor plasmid DNA with excised bacterial backbone, chemically synthesized gene fragments or enzymatically synthesized gene fragments.
The master template ensures that only target sequence is fed into our rolling circle amplification and consequently, limits the presence of impurities or unwanted sequences in the final product. This is a critical determinant of product quality.
Q: What specific challenges in plasmid-based systems do you aim to overcome with the master template?
Tejaswini: Traditional plasmid systems suffer from several "biological burdens" that we aim to eliminate with our master template synthesis, such as:
Presence of bacterial sequences: Plasmid DNA contains antibiotic resistance genes and bacterial origins of replication that often pose safety concerns in final therapeutic applications.
Target sequence ratio: The master template contains only desired target sequences, avoiding amplification of undesired sequences from the very beginning, resulting in high substrate to target DNA conversion and consequently improved yields and product costs.
Structure flexibility: The resulting master template enables replication of highly complex sequences, up to 20kb in size.
Q: If you start with a plasmid precursor, can the master template still be considered cell-free?
Tejaswini: The plasmid precursor provides the blueprint innovators are looking for in their DNA template. Through our manufacturing process, we remove the bacterial backbone from the precursor, along with any other antibiotic-resistant genes, and ORI, leaving only the gene of interest (GOI). Subsequent large scale DNA production is cell-free and early removal of these elements eliminates the contamination risks from the host-cells along with improving manufacturing timelines by removal of additional purification steps.
Fully synthetic workflows have also been validated and entirely avoid the plasmid precursor. This way the template generation can be fully decoupled from constraints of DNA manufacturing in biological host and transition towards a controlled enzymatic production system. This accelerated route is most suitable for applications such as personalized cancer immunotherapies, where time is of the essence.
Q: How does this master template impact downstream applications?
Tejaswini: The removal of bacterial sequences before DNA amplification provides major advantages for multiple downstream therapeutic applications. For instance, in both viral vector manufacturing and gene editing applications, minimizing the delivery of non-target DNA is key as those sequences can increase immunogenic response. By removing these sequences at the template generation step, our platform reduces non-specific encapsidation as well as non-specific genome integration.
While for mRNA synthesis applications, the purity of the DNA template required for in vitro transcription (IVT) is a critical factor determining quality of the mRNA product. A backbone-free template eliminates unnecessary bacterial sequences, which means there’s no risk of genomic DNA which could introduce unwanted byproducts. The use of synthetic DNA as an IVT template also means that less DNA is required to achieve equivalent yields compared to plasmid DNA.
Q: How does this approach support seamless scale-up from preclinical to commercial supply?
Tejaswini: Template generation helps reduce manufacturing risk by simplifying the production process and removing key biological variables, which is particularly important in the scale-up process. In plasmid DNA manufacturing, the process typically transitions from small shake flasks to large bioreactors, which introduces changes in oxygen transfer, nutrient availability, and shear stress that can impact product quality.
Our process is enzymatic and cell-free, making scaling much more predictable. The same reaction conditions apply across different volumes, meaning the process behaves consistently whether performed at millilitre or litre scale. This ensures seamless tech transfer to our HQ/GMP facility when producing DNA for pre-clinical and clinical studies.
Shifting to synthetic DNA also removes the need for master cell banks, including full characterization and storage, which would typically add costs and time to drug development timelines. The master template created during 4basebio’s manufacturing process is cell-free and can be used to reproduce repeat batches, ensuring both speed and consistency.
Q: What excites you most about how this platform could shape the future of genetic medicine manufacturing?
Tejaswini: One of the most exciting aspects of this platform is its potential to accelerate the development of next-generation genetic medicines. By removing the limitations imposed by bacterial systems, we can enable faster and more flexible DNA manufacturing, which is particularly important for emerging therapeutic areas such as personalized medicines.
A strong example is our collaboration with Neomatrix, where we supported them with delivering synthetic DNA for their personalized cancer vaccine (PCV) program. Using our synthetic DNA platform, it was possible to develop a personalized therapeutic candidate within a 6–8-week timeframe, demonstrating the speed and adaptability required for patient-specific treatments.
Innovations like this have the potential to reduce development timelines, improve manufacturing reliability, and ultimately enable more agile, patient-centric therapeutic approaches in areas such as oncology and gene therapy.
Tejaswini Mahadeva
Tejaswini is a molecular bio scientist with expertise in synthetic DNA technologies, gene expression systems, and molecular biotechnology. She currently serves as the Template Generation Team Lead at 4basebio, where she leads the development and optimization of template generation processes that underpin the company’s synthetic DNA production platform for gene therapy and vaccine development.
Alongside her leadership responsibilities, she contributes to research and development initiatives exploring next-generation DNA modalities and nucleic acid technologies. Her scientific background spans synthetic biology, molecular biology, protein engineering and purification, gene editing, and structural biology, reflecting a multidisciplinary approach to advancing biotechnology and nucleic acid-based therapeutics.
For more information on 4basebio’s synthetic DNA technology visit our Technology Page or if you’d like to speak to our team to discuss a project, use the button below.